Obtains predictions and corresponding credible intervals from an object of class 'JointAI'.
Arguments
- object
object inheriting from class 'JointAI'
- outcome
vector of variable names or integers identifying for which outcome(s) the prediction should be performed.
- newdata
optional new dataset for prediction. If left empty, the original data is used.
- quantiles
quantiles of the predicted distribution of the outcome
- type
the type of prediction. The default is on the scale of the linear predictor (
"link"or"lp"). Additionally, for generalized linear (mixed) models (incl. beta and log-normal)type = "response"transforms the predicted values to the scale of the response, and for ordinal and multinomial (mixed) modelstypemay be"prob"(to obtain probabilities per class),"class"to obtain the class with the highest posterior probability, or"lp". For parametric survival modelstypecan be"lp"or "response", and for proportional hazards survival models the options are"lp","risk"(=exp(lp)),"survival"or"expected"(=-log(survival)).- start
the first iteration of interest (see
window.mcmc)- end
the last iteration of interest (see
window.mcmc)- thin
thinning interval (integer; see
window.mcmc). For example,thin = 1(default) will keep the MCMC samples from all iterations;thin = 5would only keep every 5th iteration.- exclude_chains
optional vector of the index numbers of chains that should be excluded
- mess
logical; should messages be given? Default is
TRUE.- warn
logical; should warnings be given? Default is
TRUE.- return_sample
logical; should the full sample on which the summary (mean and quantiles) is calculated be returned?#'
- ...
currently not used
Value
A list with entries dat, fit and quantiles,
where fit contains the predicted values (mean over the values
calculated from the iterations of the MCMC sample), quantiles
contain the specified quantiles (by default 2.5% and 97.5%), and dat
is newdata, extended with fit and quantiles (unless
prediction for an ordinal outcome is done with type = "prob", in
which case the quantiles are an array with three dimensions and are
therefore not included in dat).
Details
A model.matrix \(X\) is created from the model formula
(currently fixed effects only) and newdata. \(X\beta\) is then
calculated for each iteration of the MCMC sample in object, i.e.,
\(X\beta\) has n.iter rows and nrow(newdata) columns. A
subset of the MCMC sample can be selected using start, end
and thin.
Note
So far,
predictcannot calculate predicted values for cases with missing values in covariates. Predicted values for such cases areNA.For repeated measures models prediction currently only uses fixed effects.
Functionality will be extended in the future.
Examples
# fit model
mod <- lm_imp(y ~ C1 + C2 + I(C2^2), data = wideDF, n.iter = 100)
# calculate the fitted values
fit <- predict(mod)
#> Warning:
#> Prediction for cases with missing covariates is not yet implemented. I
#> will report “NA” instead of predicted values for those cases.
# create dataset for prediction
newDF <- predDF(mod, vars = ~ C2)
# obtain predicted values
pred <- predict(mod, newdata = newDF)
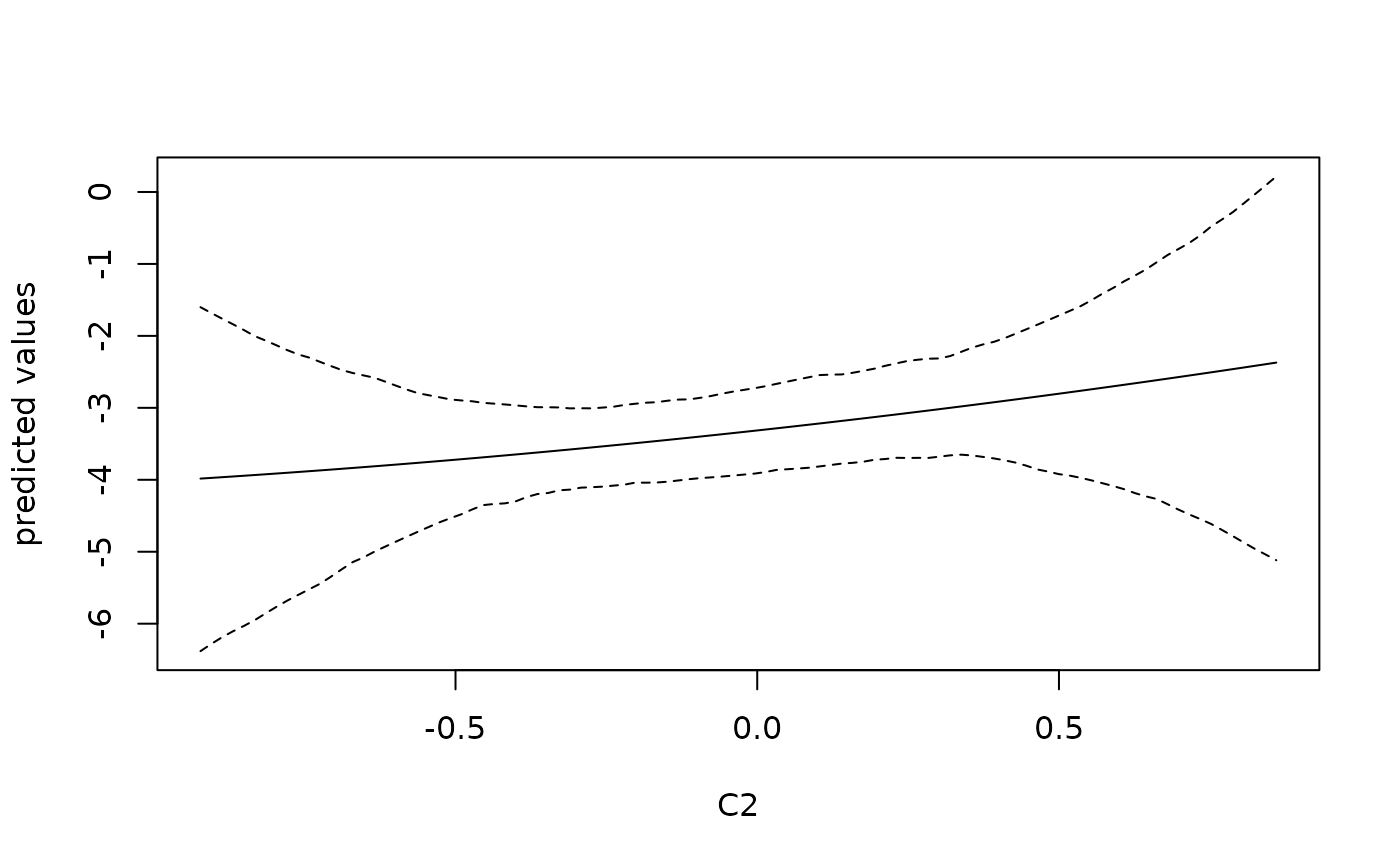
# plot predicted values and 95% confidence band
matplot(newDF$C2, pred$fitted, lty = c(1, 2, 2), type = "l", col = 1,
xlab = 'C2', ylab = 'predicted values')